Opening Vision

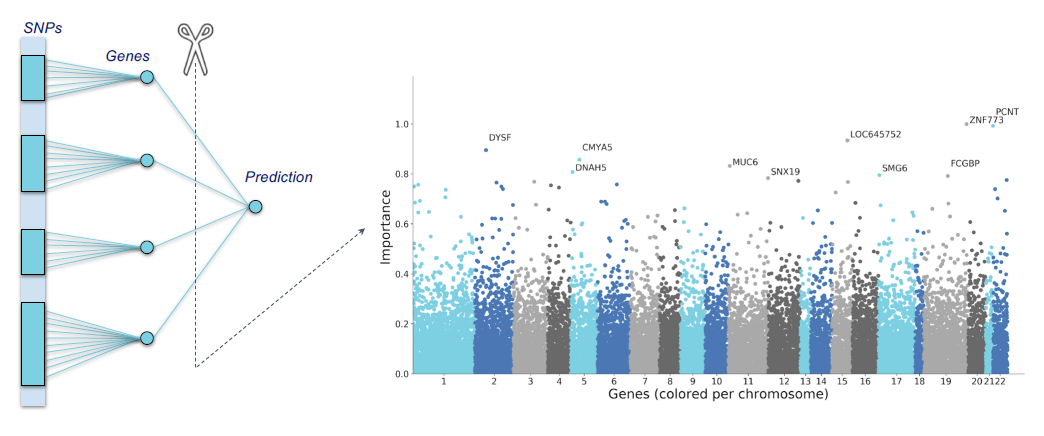
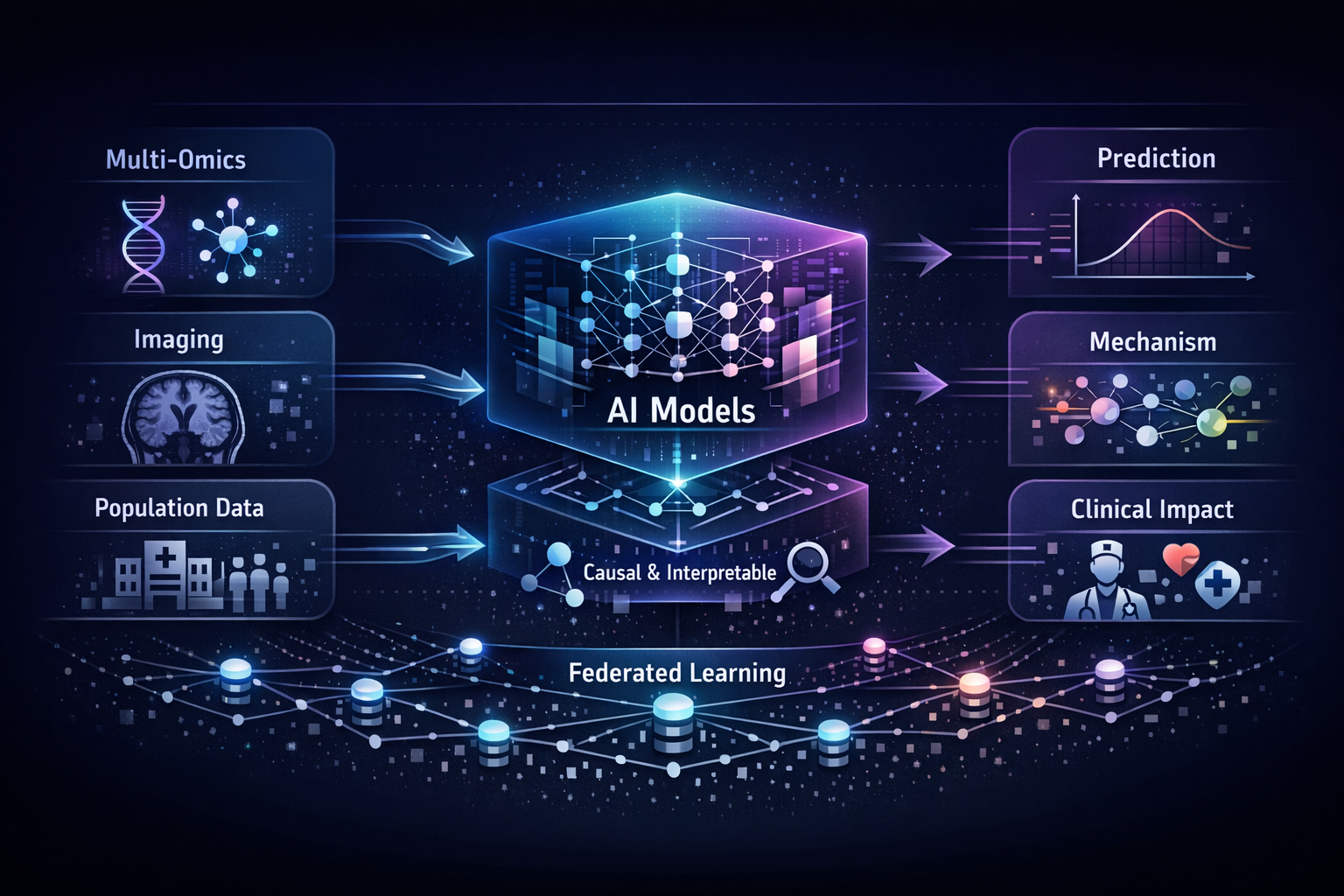
Human health is written across genomes, images, and lifetimes of exposure and care. In our group, we study how these layers fit together in large population cohorts such as the Rotterdam Study and Generation R, and how they can be translated into models that are both predictive and biologically meaningful. We connect molecular variation to organ-level structure and function, and population context to risk, using integrative methods that combine multi-omics, quantitative imaging, and longitudinal clinical and lifestyle data with international consortia. The aim is not prediction alone, but a clearer path from population evidence to mechanism, and from mechanism to prevention and precision medicine.
The group consists of PhD candidates, postdoctoral researchers, and research engineers working at the interface of epidemiology, AI, and imaging. Projects are typically embedded in large cohort studies and international collaborations, combining methodological development with applied research.
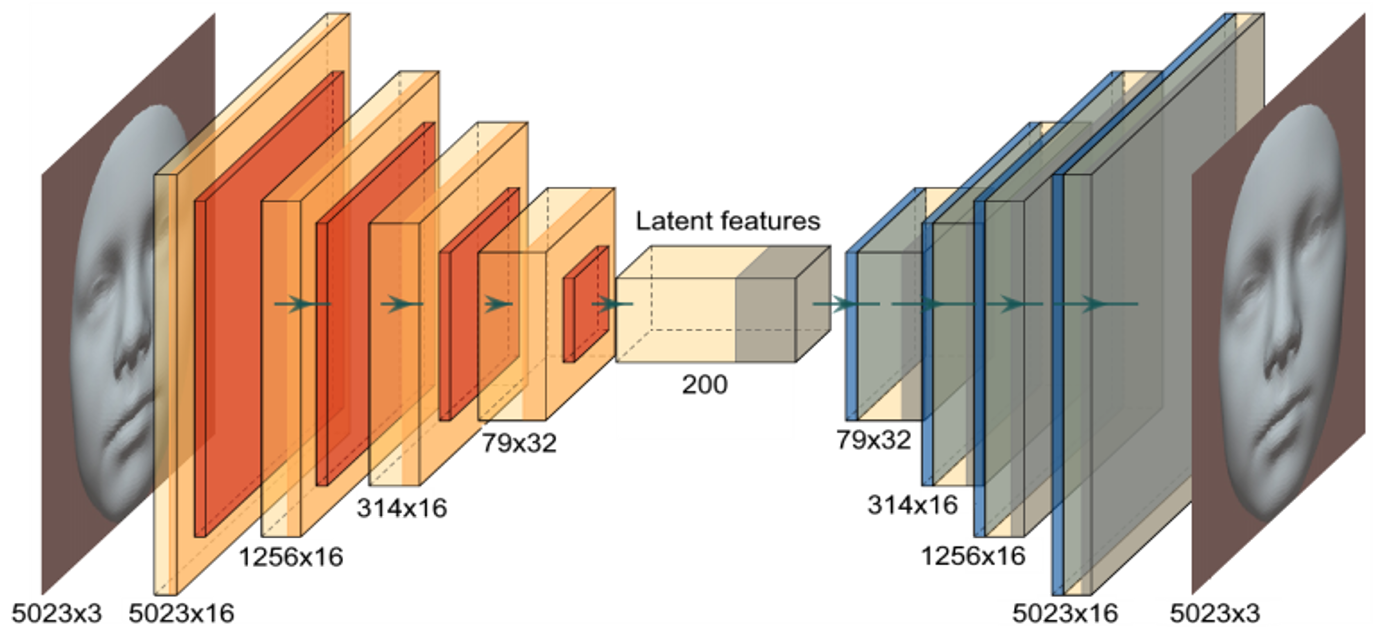
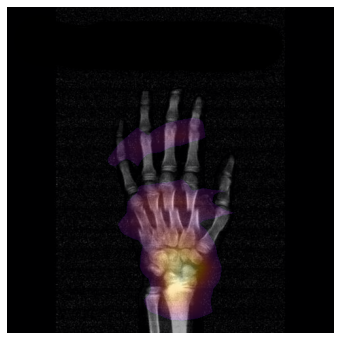
Imaging and computation
Embedded collaboration with the Biomedical Imaging Group Rotterdam (BIGR): large-scale imaging science, GPU computing, and infrastructure for AI-driven discovery.
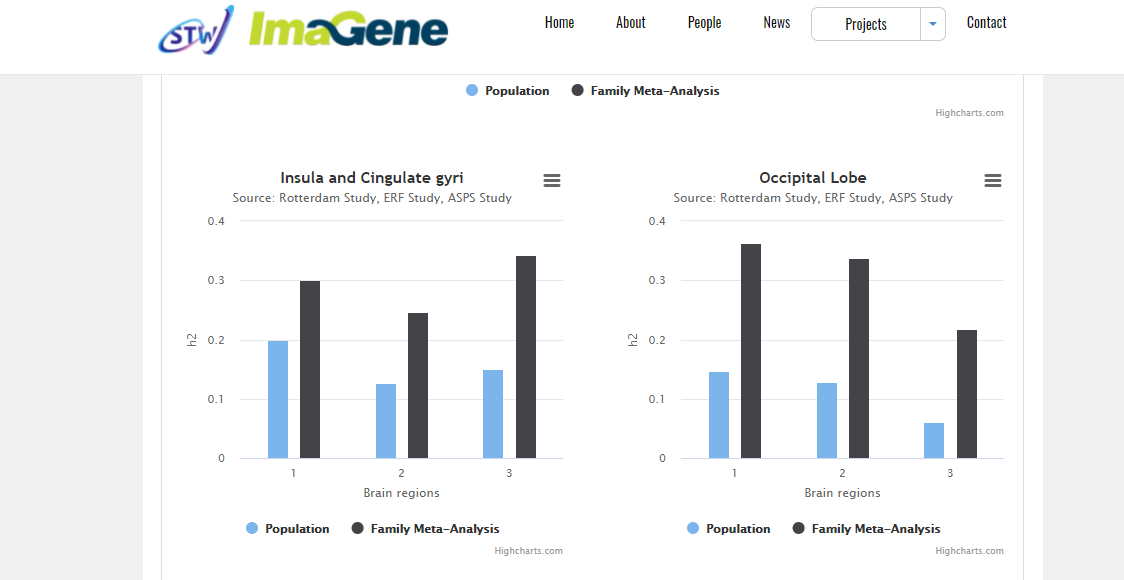
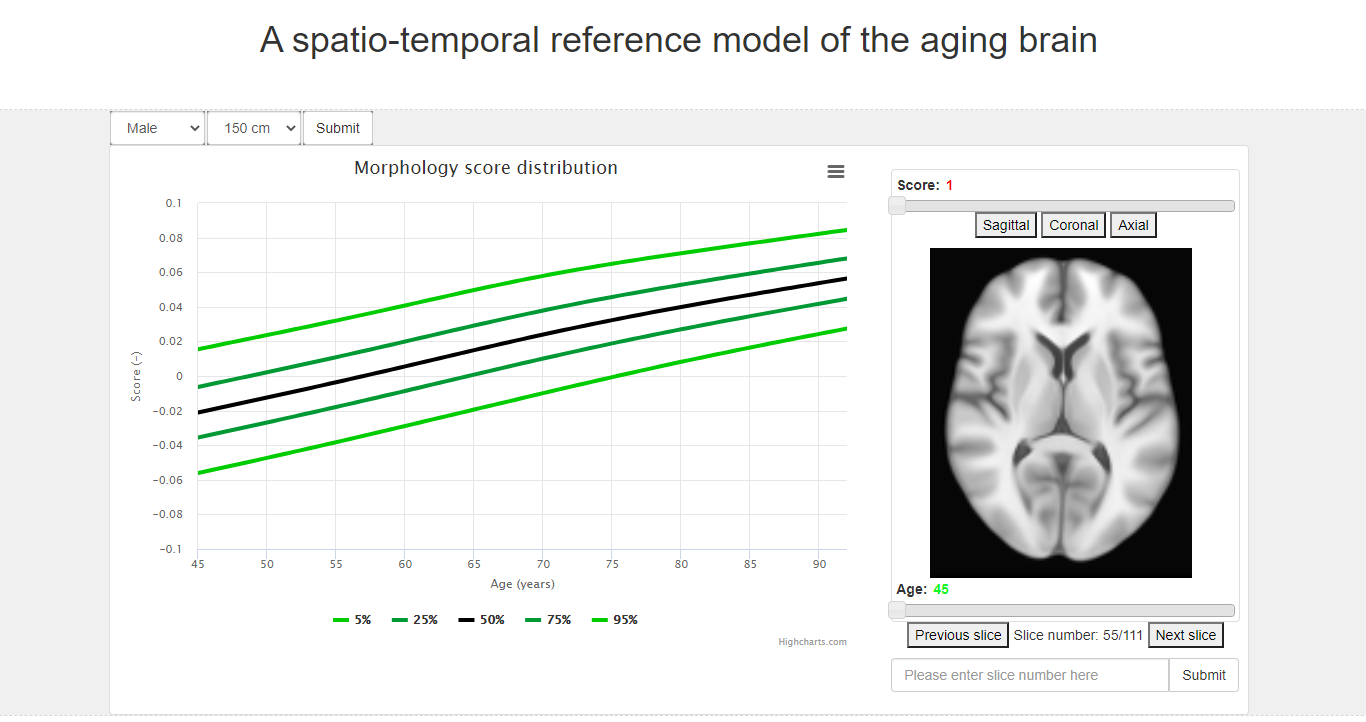
Population cohort depth
Prospective resources including the Rotterdam Study (ERGO) and training through NIHES, extended via meta-analysis and international harmonization.